Abstract
Background
Multiple Myeloma (MM) is an incurable plasma cell (PC) malignancy and high risk MM remains an unmet clinical need. Translocation 4;14 occurs in 15% of MM and is associated with an adverse prognosis. A deeper understanding of the biology and immune micro-environment of t(4;14) MM is necessary for the development of effective targeted therapies. Single Cell multi-omics provides a new tool for phenotypic characterization of MM. Here we used Proteona's ESCAPE™ single cell multi-omics platform to study a cohort of patients with t(4;14) MM.
Methods
Diagnostic bone marrow (BM) samples from 13 patients with t(4;14) MM (one of whom had samples at diagnosis and relapse) were analysed using the ESCAPE™ platform from Proteona which simultaneously measures gene and cell surface protein expression of 65 proteins in single cells. Cryopreserved BM samples were stained with antibodies and subsequently sorted on CD138 expression. The CD138 positive and negative fractions were recombined at a 1:1 ratio for analysis using the 10x Genomics 3' RNAseq kit. Resulting data were analyzed with Proteona's MapSuite™ single cell analytics platform. In particular, Mapcell was used to annotate the cells and MapBatch was used for batch normalization in order to preserve rare cell populations.
Results
Patients had a median age of 63 years and received novel agent-based induction. Median progression free and overall survival (PFS and OS) were 22 and 34 months respectively.
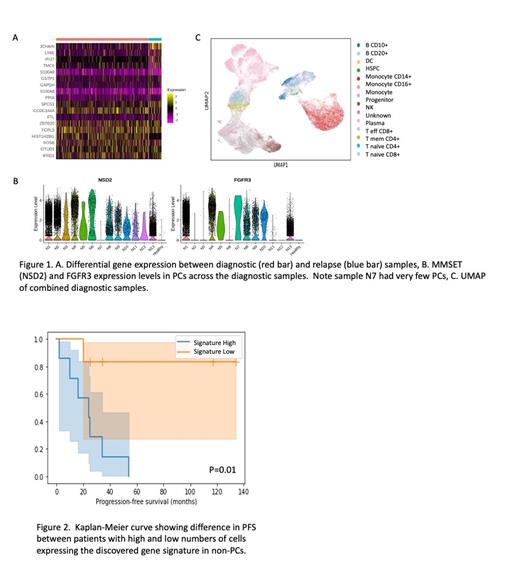
We first analyzed serial BM samples from an individual patient that were taken at diagnosis and relapse following bortezomib based treatment. The PCs in this patient showed variations in gene expression between diagnosis and relapse (Fig 1A), including the reduction of HIST1H2BG expression, which has previously been correlated with resistance to bortezomib. Subsequent analysis of the immune cells identified a shift in the ratio of T cells to CD14 monocytes from 5.7 at diagnosis to 0.6 at relapse suggesting a major change in the BM immune micro-environment in response to therapy.
Next, we analyzed the malignant PCs of the diagnostic samples. As expected, MMSET (NSD2) was overexpressed in all PCs compared to normal PCs, while FGFR3 expression could be categorized into no expression of FGFR3, low expression (<10% of cells expressing FGFR3) or high expression (>80% of cells expressing FGFR3) (Fig 1B). No gene or protein expression patterns within the PCs were identified that correlated with PFS or OS in this cohort.
Finally, we analyzed the immune micro-environment in the diagnostic samples (Fig 1C). While there was no overall discernable pattern of cell types present, one cluster of cells, annotated as 'unknown' cell type, suggested a small population of cells that had not been previously annotated in published single cell RNA-seq data. The cells were CD45+ and CD138 - both at the protein and RNA level, suggesting they are not plasma cells.
We tested if the number of the 'unknown' cells in each sample correlated with PFS, but there was no significant correlation. We then used these cells to derive a gene signature profile which was expressed in most of the cells in the 'unknown' cluster as well as a minor fraction of cells in other clusters including some PCs. The number of cells expressing the gene signature negatively correlated with PFS, with samples containing more cells expressing the signature having a lower PFS than samples with fewer signature positive cells (Fig 2). The correlation remained significant whether we included PCs in the analysis or not, but was not significant amongst only the PC population, suggesting that the cells responsible for the correlation are from the immune micro-environment.
Conclusions
We present the first application of single cell multi-omic immune profiling in high-risk MM and demonstrate that t(4;14) is a phenotypically heterogenous disease. While no consistent gene or protein expression patterns were identified within the malignant cell population, we did identify gene expression changes in a relapsed patient sample that may reflect key alterations in the PCs responsible for therapy resistance. In addition, we identified a gene signature expressed in a rare population of non-plasma cells that significantly correlated with PFS in this patient cohort. These data highlight the potential of single cell multi-omic analysis to identify immune micro-environmental signatures that correlate with response to therapy in t(4;14) MM.
Scolnick: Proteona Pte Ltd: Current holder of individual stocks in a privately-held company. Huo: Proteona Pte Ltd: Ended employment in the past 24 months. Xu: Proteona Pte Ltd: Current Employment. Chng: Amgen: Honoraria, Research Funding; Janssen: Honoraria, Research Funding; Celgene: Honoraria, Research Funding; Novartis: Honoraria; Abbvie: Honoraria.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal